This will allow you to go back to the IPA preferences and set it back to the original memory setting that had worked on your computer.ĭisplaying the pre-loaded Example Analyses can be disabled under Projects. If this happens, please go to this URL and start IPA with this URL. If you create large Comparison Analyses, want to use more advanced features (such as Causal Network Analysis, IsoProfiler, Analysis Match, etc.), set IPA to >4000 Mb if you have enough installed RAM on your computer. However, note that IPA won't start up if you allocate it more memory than is available in your system memory. If you have a computer with 6 GB or more, set IPA to 2000 or more. Ideally, if you have at least 4 GB of installed RAM on your computer, you would set the setting to at least 1000 or 1200 mb. 
IPA needs at least 750 Mb of system memory to be available to it. The amount of system memory (i.e., the amount of RAM) allocated for the IPA application can be set using the pull-down menu. For the observation you would like to change, click on the colored square, then choose a color for the color palate. You can change the colors of the bars in Canonical Pathways, Diseases & Functions Analysis, My Pathways Analysis, and List Analysis. Clicking on a colored square will open the color selector, from which you can select the color of your choosing. You must restart IPA before any changes go into effect.ĪPPLICATION tab of Application Preferences:Īllows you to customize the colors that signify up-and-down-regulation for the Expression Value types. You can click on the "Restore Defaults" button to revert to the original settings. The application's preferences can be accessed from the drop-down menu by selecting File->Preferences->Application Preferences.Īll options require that any modification to a preference be retained only if you click on the Save button at the bottom of the relevant tab. To access these videos, click here.Change key settings for IPA's memory usage, overlay colors, etc. Finally, Qiagen hosts webinars addressing the use of and updates to this software. 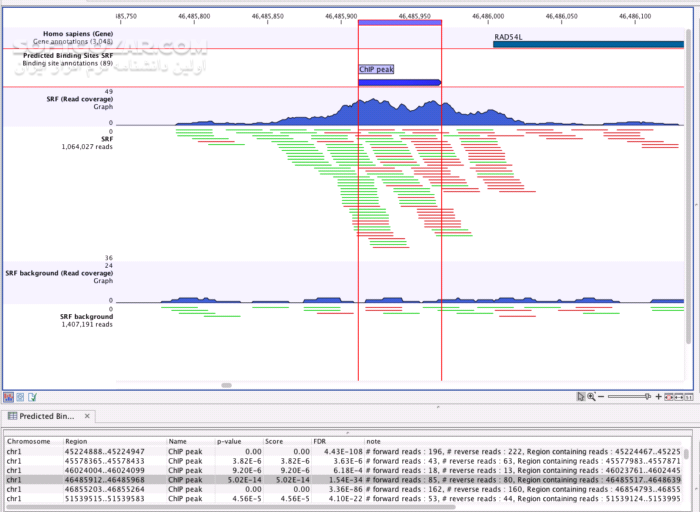
Additionally, there are tutorials available for different workflows. For the CLC Genomics Workbench manual, click here. Getting Helpĭocumentation for the CLC Genomics Workbench is available under the Help tab in the software. Working with CLC Genomics Workbench requires login to the NIH network or VPN connection if remote. This software requires access to a floating license server (three simultaneous users), and so care should be taken to return the license when the software is not actively being used (i.e. You must submit a request through to obtain access to CLC Genomics Workbench.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
